-
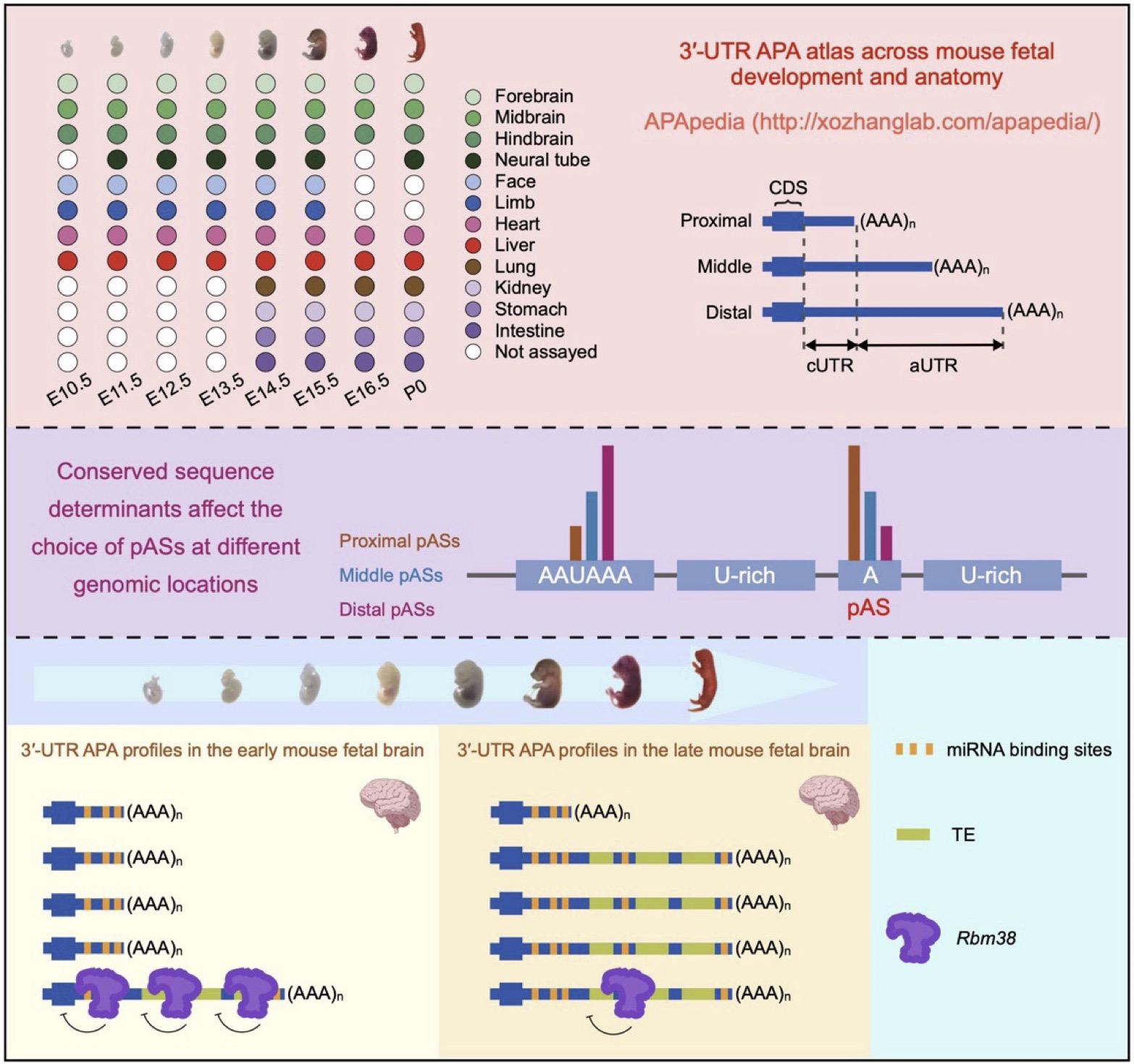
APApedia database
APApedia is a comprehensive database dedicated to unraveling the dynamics of alternative cleavage and polyadenylation (APA) during developmental processes. It provides an extensive collection of APA events in humans and mice, encompassing a wide range of developmental contexts. (Wang et al., Advanced Science, 2025)
-
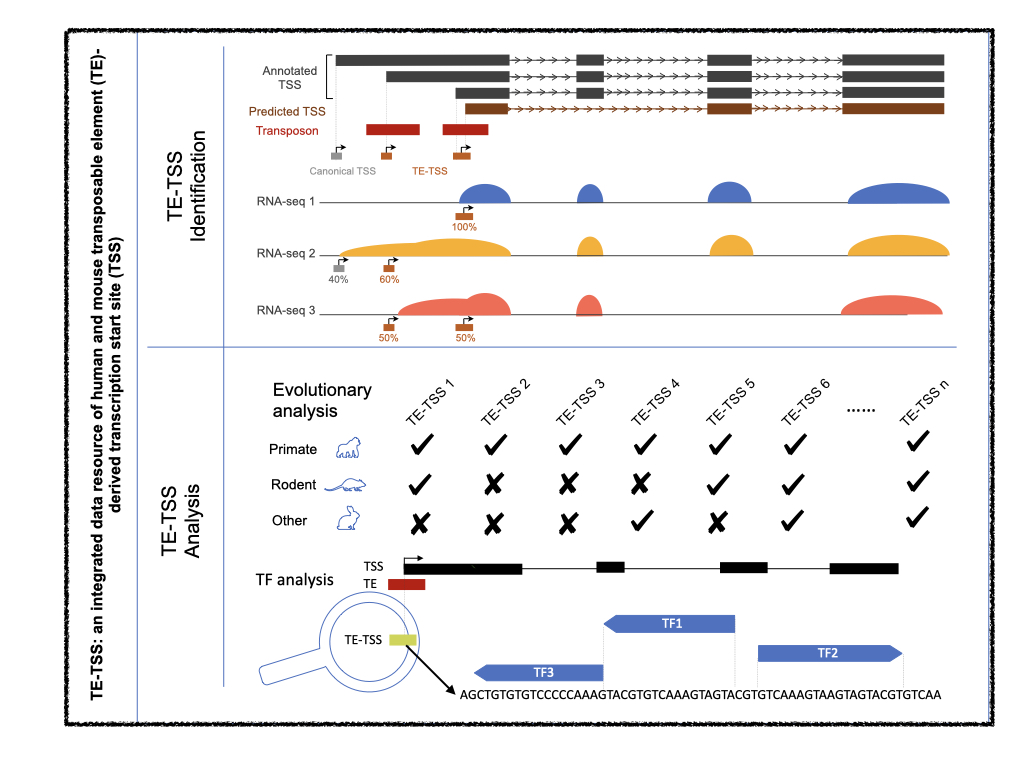
TE-TSS database
TE-TSS is a database for transposon (TE)-derived transcription start sites (TSSs). It provides a comprehensive collection of TE-derived TSSs in humans and mice, and enables users to search, browse, and download TETSSs and their usage across different biosamples. (Gu et al., Nucleic Acids Research, 2023)
-

RAMPAGE analysis toolkit
RAMPAGE analysis toolkit is a standardized pipeline designed to execute quality control, data cleaning, peak calling, and peak annotations for RAMPAGE data. Using this pipeline, we successful identify Pol III-transcribed Alu elements, alternative TSSs, expressed snRNA variants, and transposon-derived TSSs. (Zhang et al., Genome Research, 2019) (Moore et al., Genome Research, 2022)
-
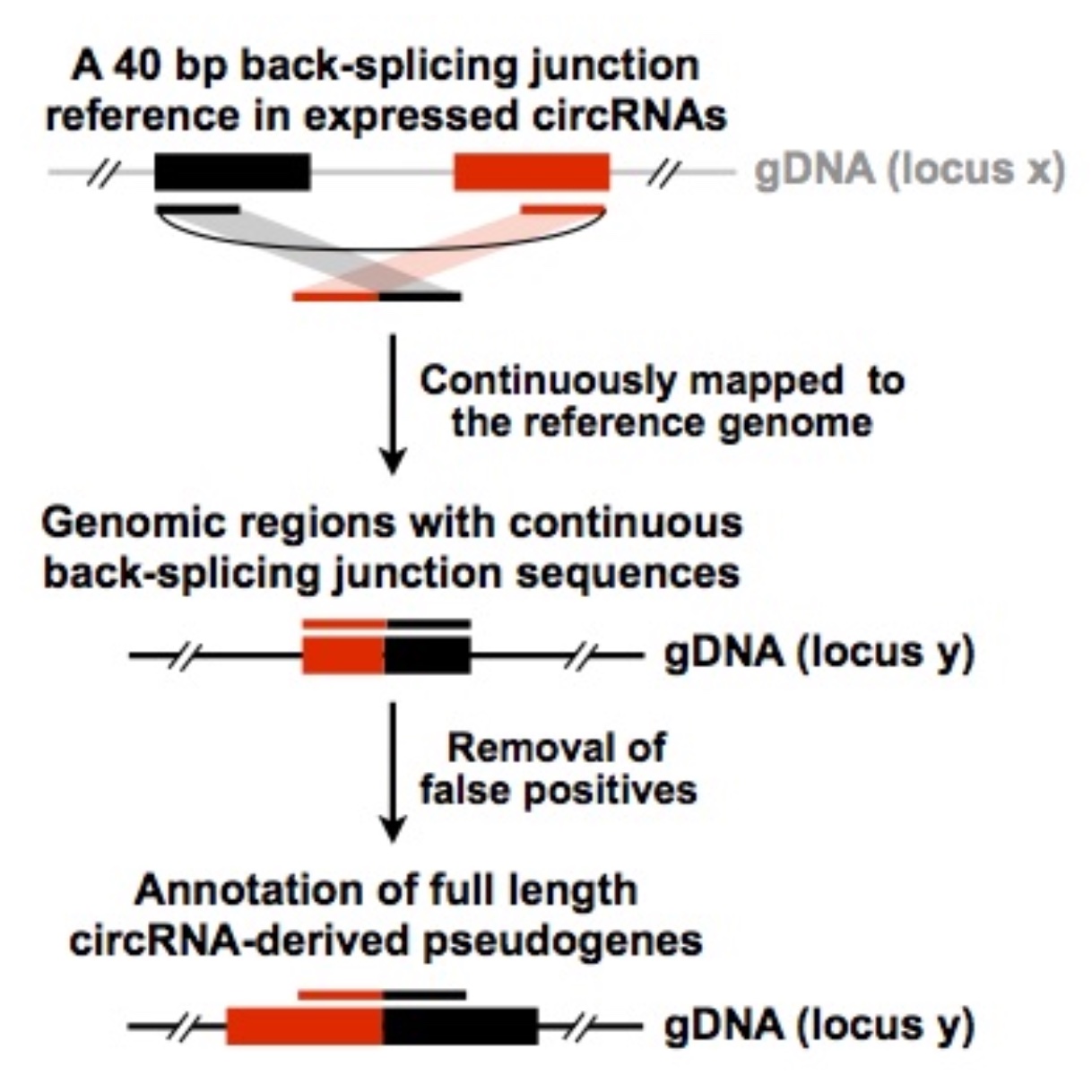
CIRCpseudo
CIRCpseudo is a pipeline to map back-splicing junction sequences for circRNA-derived pseudogenes. Using this pipeline, you could identify potental pseudogenes derived from circRNAs. (Dong et al., Cell Research, 2016)
-

CIRCpedia
CIRCpedia is an integrative database, aiming to annotating alternative back-splicing and alternative splicing in circRNAs across different cell lines. (Zhang et al., Genome Research, 2016)
-
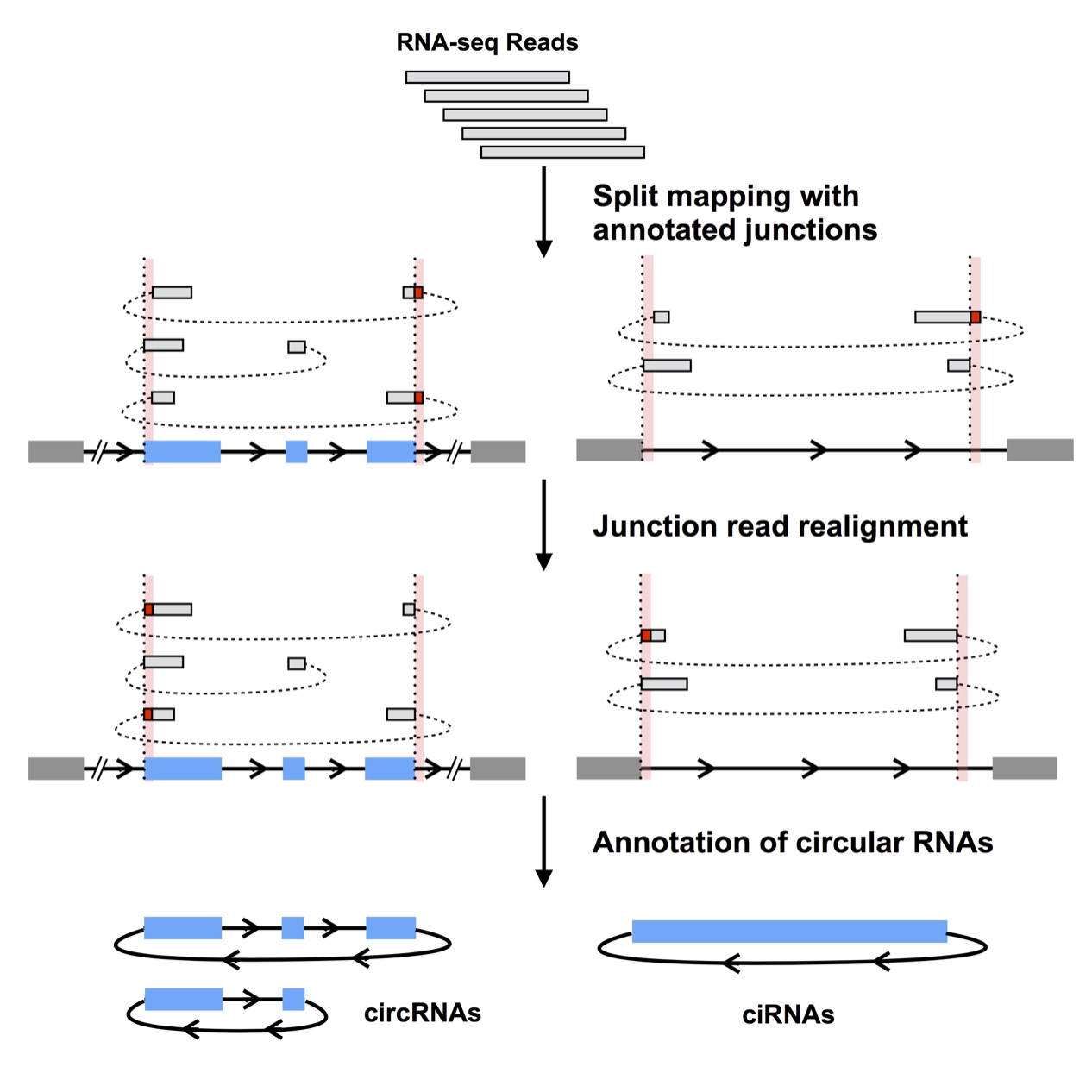
CIRCexplorer/CIRCexplorer2
CIRCexplorer is a comprehensive and integrative circular RNA analysis toolset. It offers support for multiple circular RNA aligners, enables de novo assembly of novel circular RNA transcripts, and facilitates the characterization of various alternative (back-)splicing events in circular RNAs. (Zhang et al., Cell, 2014) (Zhang et al., Genome Research, 2016)
-
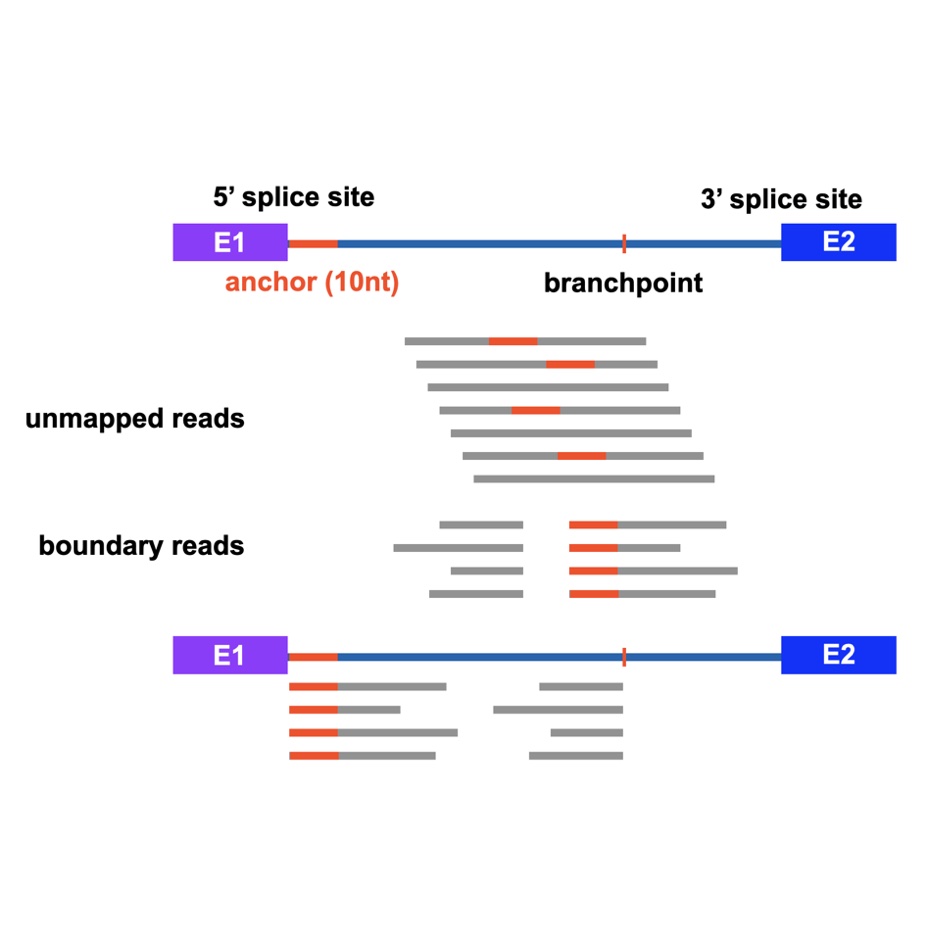
CIRCfinder
CIRCfinder is a pipeline to map junction reads for circular intronic RNAs (ciRNAs). Using this pipeline, you could determine the exact boundaries of interested ciRNAs. (Zhang et al., Molecular Cell, 2013)
Xiao-Ou Zhang Lab
Decoding noncoding sequences!