Welcome!
We are a computational biology group based at the School of Life Sciences and Technology, Tongji University. Our primary focus is to unravel the intricate
and diverse mechanisms that underlie gene regulation driven by non-coding sequences at the transcriptome
level. We specialize in the developing of sophisticated methods tailored for the statistical and integrative
analysis of multi-omics high-throughput data. Through these meticulously crafted investigative approachs, we
delve into the pivotal role of repetitive sequences, with particular emphasis on transposable elements
(TEs), in the control of gene expression. In parallel, We strive to unravel the complexity of
transcriptional and post-transcriptional gene expression regulation, along with elucidating the mechanisms
governing the biogenesis of non-coding RNAs and their vital biological functions. Our research contributes
to advancing the understanding of non-coding sequences and their relevance in human health and disease
therapeutics.
Decipher TE-Mediated Gene Expression
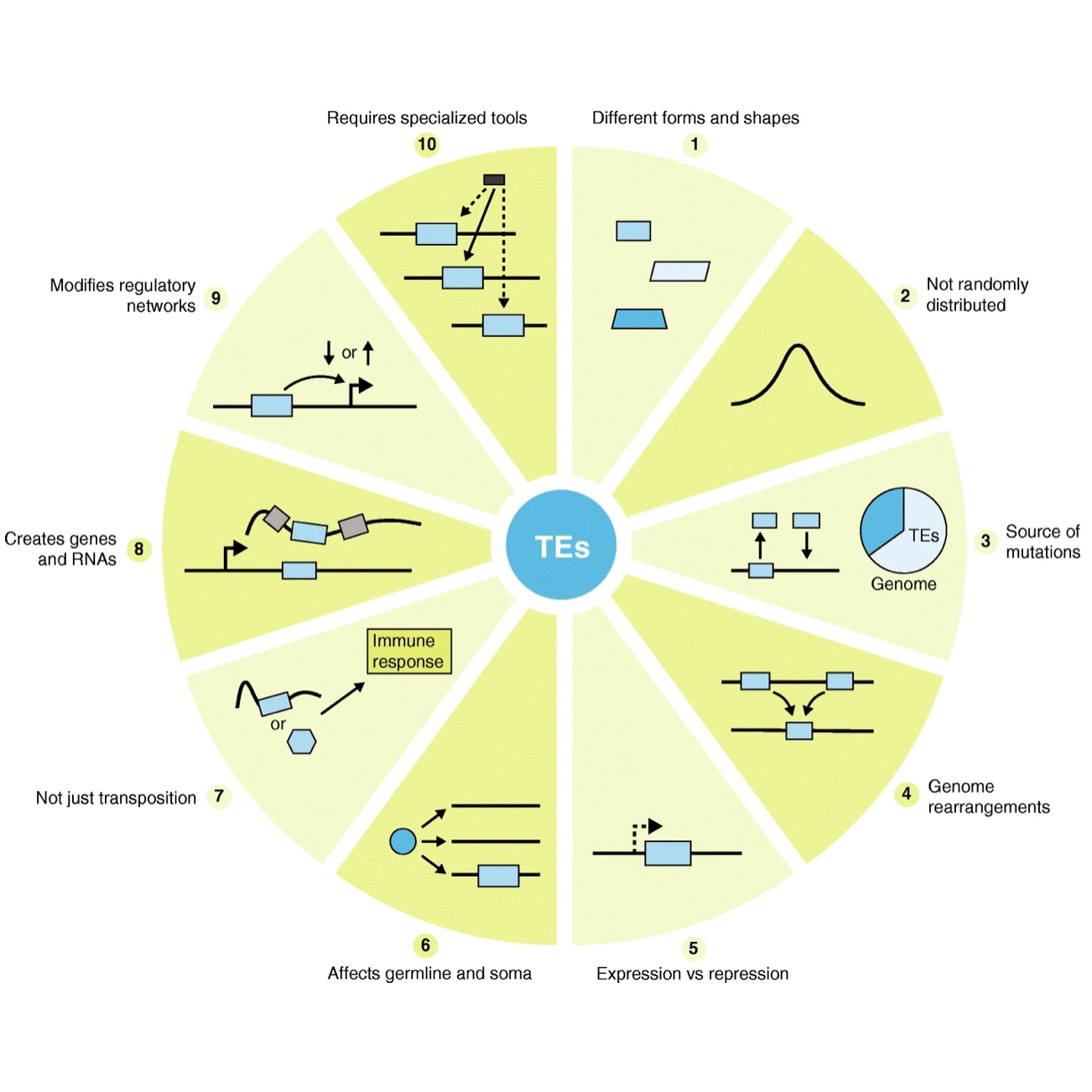
One of the primary research focuses in our lab revolves around investigating the intricate mechanisms underlying the involvement of transposable elements (TEs) in the regulation of gene expression. TEs, unique components within the genome, play a pivotal role not only in driving genome evolution but also in influencing fundamental aspects of biology and disease. Within this realm of inquiry, our efforts are dedicated to uncovering how TEs impact gene expression. Many TEs contain transcriptional regulatory elements, adding a layer of complexity to their functional diversity. Our research aims not only to reveal the mechanisms underlying the generation of new transcripts from these elements but also to unravel their roles in cellular functions and biological processes. Beyond serving as sources of new transcription, TEs can function as cis-regulatory elements. We will investigate how they interact with transcription factors, thereby influencing gene activity and expression levels. Moreover, we will explore how TEs operate within specific cellular contexts, taking into account temporal and spatial factors, and examining their connections with epigenetic modifications. To address this research direction comprehensively, we will employ a multidisciplinary approach that encompasses both experimental and computational methods. Our aim is to uncover the intricate mechanisms through which TEs contribute to the regulation of gene expression, thereby shedding new light on genome function and evolution.
Unraveling Kinetics of RNA Processing
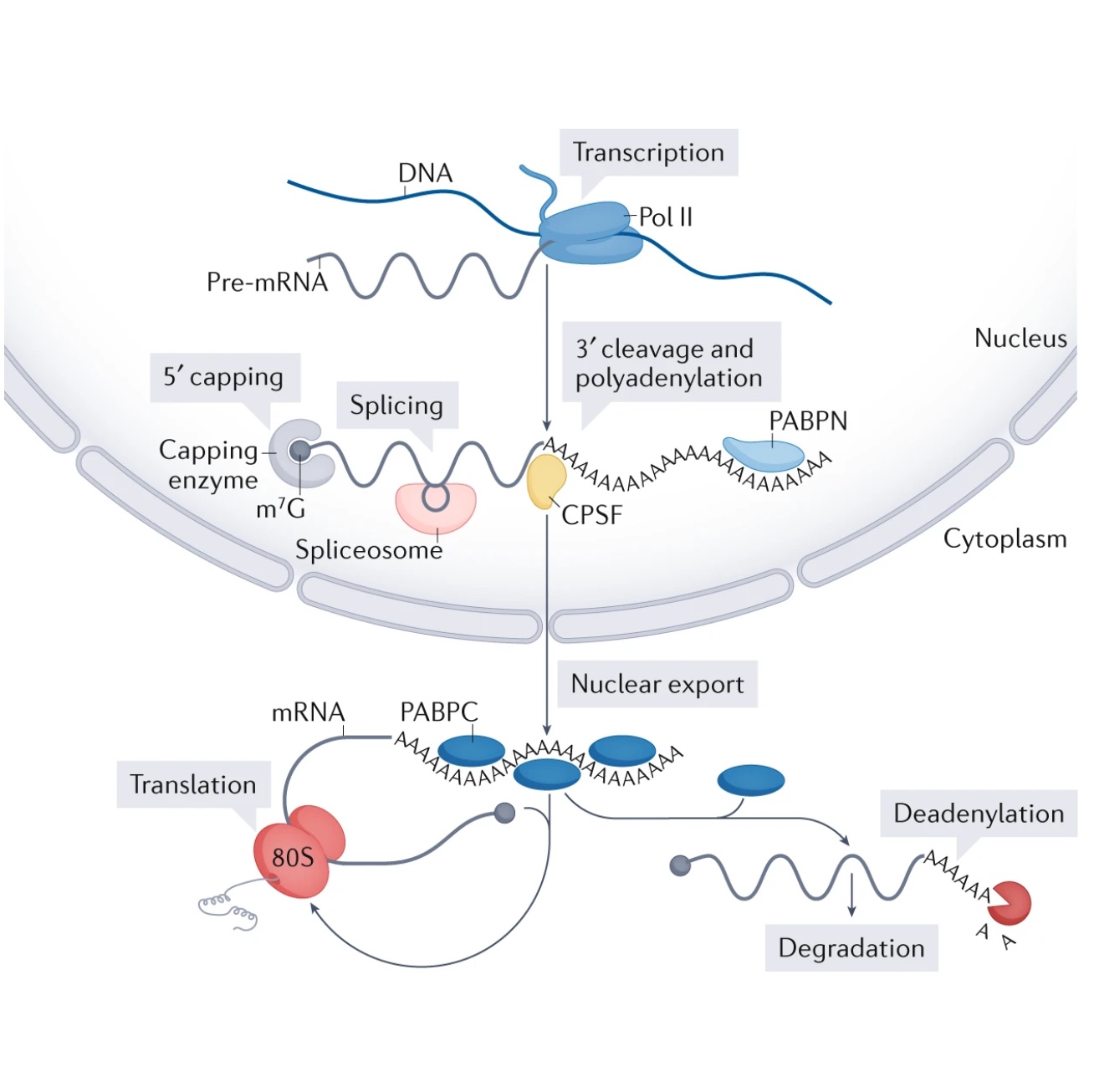
Our lab is dedicated to delving into the intricacies of RNA processing, a critical process that significantly impacts the diversity of gene expression in living organisms. Phenotypic variations in humans arise from nuanced changes in gene regulation, resulting in a wide array of gene expression patterns from the same genetic code. Recent research has provided unprecedented insights into the individual molecular mechanisms orchestrating shifts in mRNA levels, revealing that the remarkable diversity of the transcriptome stems from distinct ways RNA molecules are processed.While conventional gene expression studies often concentrate on fixed RNA levels, living systems inherently possess dynamism. This leads us to a fundamental question in functional genomics: How do the myriad dynamic molecular processes converge to shape the cellular RNA landscape? Our mission is to understand the velocity and efficiency of pivotal milestones in the RNA life cycle: the elongation phase during transcription, the intricate RNA splicing process, and the precise cleavage and polyadenylation steps. To achieve this, we employ innovative experimental techniques to capture nascent RNA molecules, followed by cutting-edge high-throughput sequencing. Complemented by computational and mathematical modeling strategies, we can simultaneously capture the rhythm of each RNA processing step along the gene. These comprehensive genome-wide measurements provide insights into the underlying principles and governing logic that dictate efficient RNA processing within cells.
Decoding Noncoding RNA Complexity
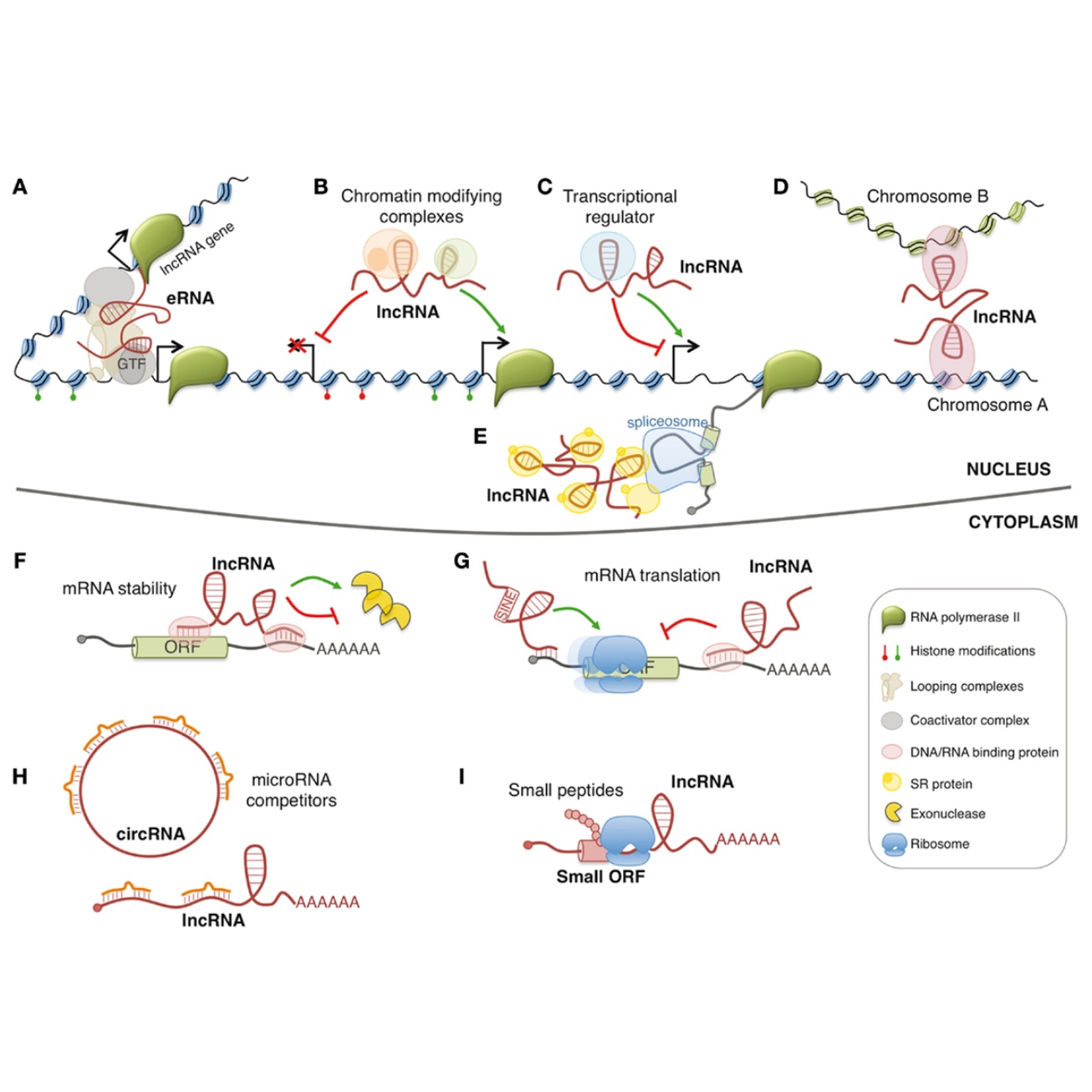
A significant proportion of human transcripts do not possess the coding potential for proteins, classifying them as noncoding RNAs. These molecules exhibit distinct expression patterns specific to various diseases, and their involvement in both transcriptional and post-transcriptional regulations positions them as potential drivers of disease. Our primary objective is to conduct an in-depth exploration of the biogenesis and molecular functions of noncoding RNAs within the intricate landscape of genetics and disease, ultimately contributing to the advancement of precision medicine. To attain this objective, we have embraced an innovative methodology that harnesses the power of mathematical and statistical modeling, combined with cutting-edge experimental techniques. This strategic combination enables us to unravel the intricate attributes of long noncoding RNAs characterized by unique structures such as circular RNAs (circRNAs) and small noncoding RNAs like snRNAs,tsRNAs, and rsRNAs. Through our comprehensive research, we strive to uncover the subtle intricacies of noncoding RNAs, thus shedding light on their potential impacts on human health and disease.


